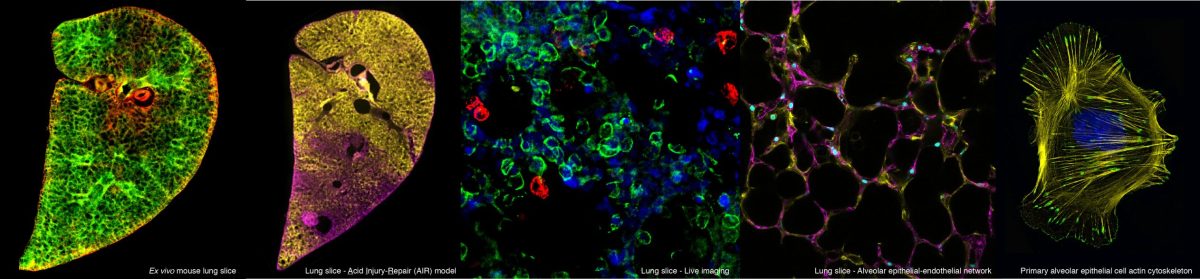
Role of developmental genes in adult lung disease and tissue repair
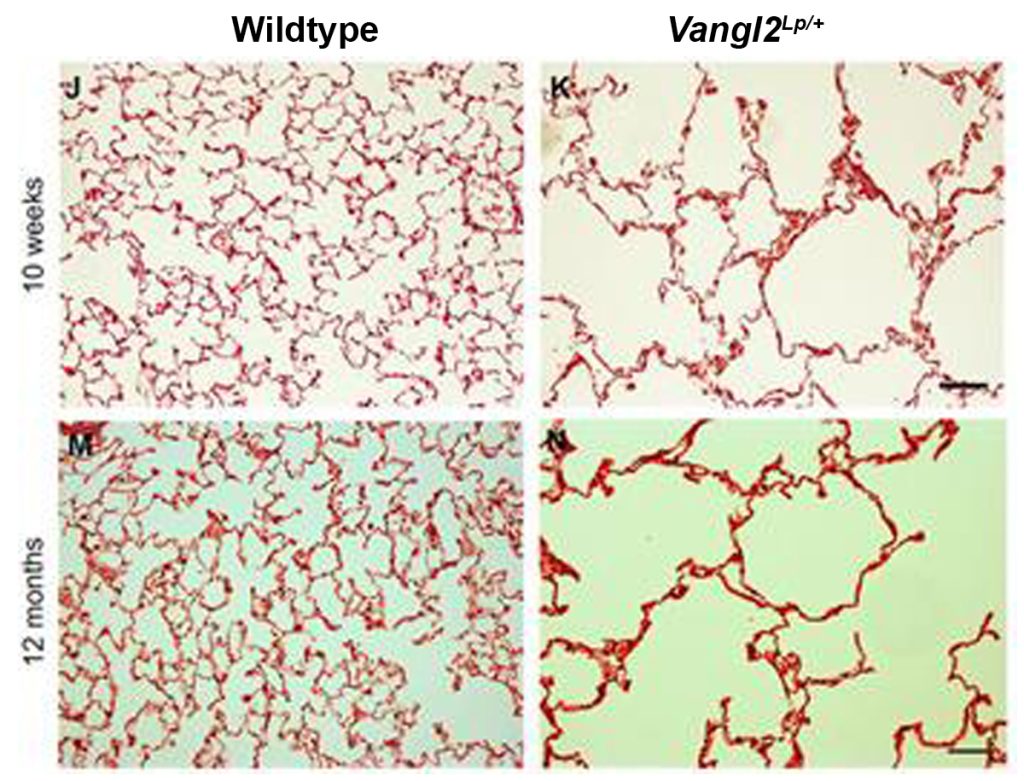
Our recent work on the Wnt/PCP pathway has shown that this pathway also plays a key role in adult lung homeostasis and repair (Poobalasingam et al. 2017 & Cheong et al. 2020)
1) We are investigating how we can manipulate the PCP pathway to modify lung repair.
2) We also study the function of other genes that we have shown are important for lung development in adult lung homeostasis and disease e.g. Wnt5a.
3) We are working on elucidating the most effective ‘lung repair cocktail’ .
Imaging lung development, injury and repair
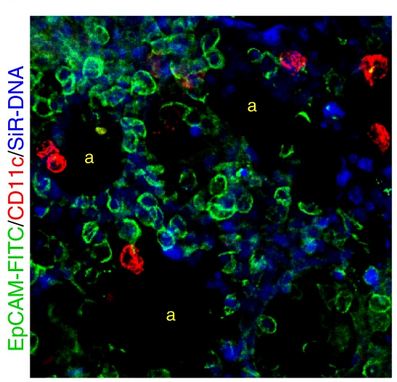
We have previously used real-time imaging to enable us to discover details about the role of genes in lung development that would not have been possible using 2D/static imaging techniques. We have established Precision-Cut Lung Slice models from human and mouse tissue, future work will continue to use this important pre-clinical tool.
1) Real-time modelling of lung injury in slices
We use this system to assess potential novel treatments to manipulate lung repair.
 Click here to view the experimental flow of the application of Acid Injury and Repair (AIR) model
Click here to view the experimental flow of the application of Acid Injury and Repair (AIR) model
2) Stretch of lung slices
We are exploring this model for studies in lung mechanobiology.
3) Real-time imaging of alveolarisation
Alveoli, the gas-exchanging compartment of the lungs, are thought to form by cells repeatedly sub-dividing airspaces eventually creating the large surface area needed for respiration. Despite their critical function, current knowledge is based solely on 2D pictures. We have used live 3D imaging of alveolarisation in slices to determine precisely how alveoli form (Akram et al. 2019).
 Click here to view movies generated from real-time imaging of lung slices.
Click here to view movies generated from real-time imaging of lung slices.
Genetics of lung disease
Working with specialists in population health (epidemiologists and medical statisticians) we are combining population genetic data, for example from UK biobank, with laboratory investigations to identify and understand genetic causes of lung disease (Portas and Pereira et al. 2020).
Key collaborators: Dr Matthew Hind, Dr Cosetta Minelli, Prof. Peter Burney, and Prof Seif Shaheen
Mouse models of lung disease susceptibility genes
We are investigating both point mutants and null mutants of human lung disease associated genes in the mouse. This work will enable us to confirm the identity of candidate disease susceptibility genes identified in human studies and to understand the function of these genes.
Lung-derived extracellular vesicles: how they are altered in response to ageing
We are investigating how the physicochemical and biological composition of extracellular vesicles (EVs) are altered in the ageing lung. EVs are heterogeneous particles (50 – 1000 nm) secreted by cells that share biological information between cells through encapsulated signalling proteins, nucleic acids and lipids. Research into EV biology has grown exponentially since their potential as non-cellular tissue modifiers has begun to be realised. We are currently addressing a critical gap in the knowledge about EV biology using a variety of tools and techniques previously established in the lab: precision-cut lung slices, the acid injury and repair (AIR) model, fluorescence microscopy, and atomic force microscopy.